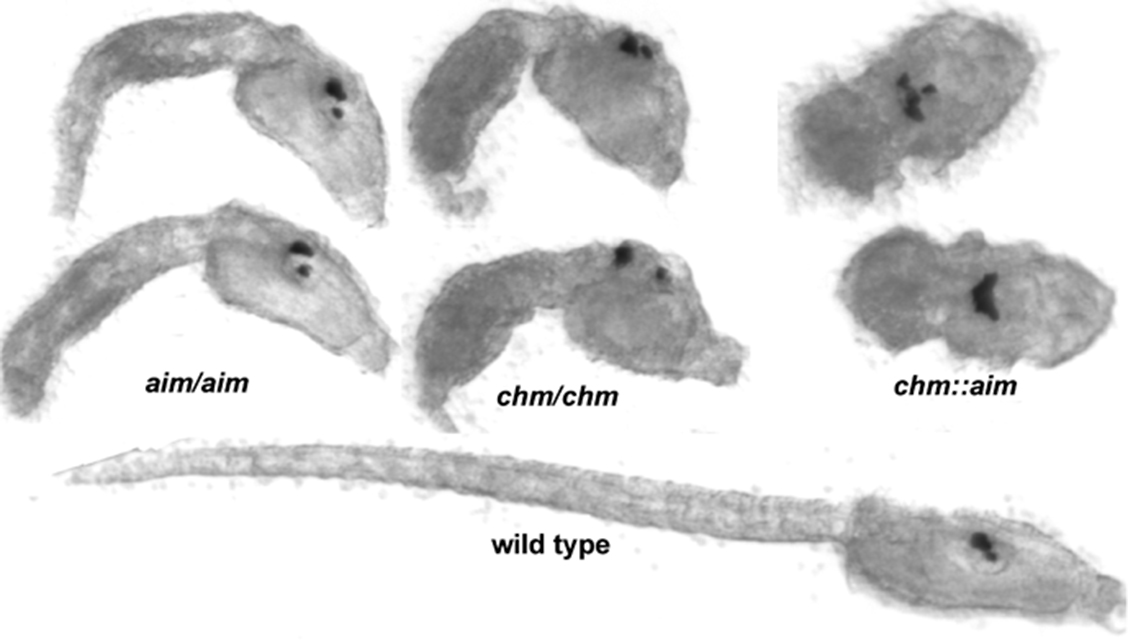
Isolation of Developmental Mutants in Ciona
In addition its the embryonic and genomic simplicity, Ciona has one additional feature that makes it ideal for forward genetics: they are hermaphrodites with a capacity for self-fertilization, which allows for rapid screening for recessive mutations. We have found that the wild population of Ciona has an extraordinarily high level of genomic variation, including a high load of lethal recessive mutations. By screening the wild population we have uncovered mutations disrupting major developmental pathways, and which have profound effects on the development of the CNS and notochord, as well as other organs and tissues. We are able to maintain mutants either as living stocks at the UCSB marine lab, or as cryopreserved sperm.
We have developed a rapid deep sequencing-based method for mapping mutations that replaces our old, and very slow, SNP-based mutation mapping method. In this new method, an outcrossed heterozygous parent carrying the recessive mutation is self-fertilized. DNA is isolated from a pool of ≈300 of the homozygous mutant progeny and deep-sequenced at >20X coverage. The deep sequencing reads are then aligned to the reference genome. The mutant locus is evident as a peak in homozygosity within the aligned sequencing reads. With this method we have been able to map mutations to a single gene.



