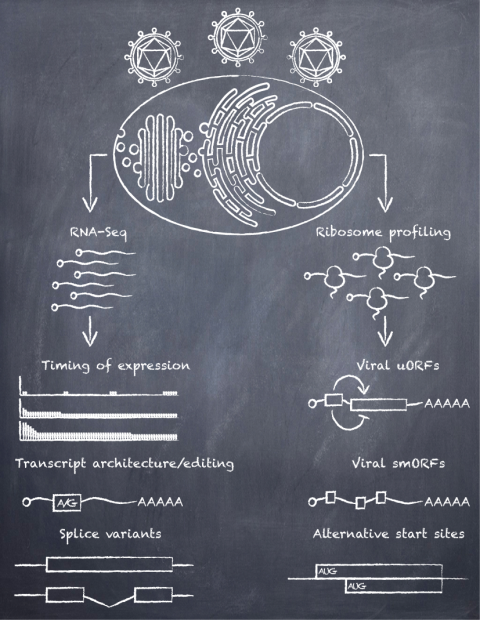
Functions of viral small open reading frames
Viral genomes encode numerous small open reading frames of unknown function. What is their importance?
The need for coordinated protein synthesis during viral infection inspired us to investigate the timing of translation, and the identity of viral features expressed during latent and lytic infection with KSHV. Ribosome profiling and RNA-seq in cells infected with KSHV (1), enabled us to characterize the detailed timing of expression of the viral transcripts and open reading frames, identify strategies used by KSHV to expand its coding repertoire, and identification of new putative coding regions. A fascinating observation emerged; the viral genome is peppered with dozens of regions that harbor short open reading frames (ORFs). These putative open reading frames (ORFs) are characterized into two categories: independently localized small ORFS (smORFs), of if they precede a primary ORF, they are known as upstream ORFs (uORFs). smORFs and uORFs have been found in other viruses, yeast, insect, plant, and mammalian cells, suggesting ancient biological roles. Their functions have been documented only in a handful of systems, and for the most part they remain a mystery. Upstream ORFs are regulatory elements that can modulate the translation of the main ORF they precede. Small ORFs can encode for functional peptides implicated in diverse processes. The relatively small size of its genome, and the discrete numbers of smORFs and uORFs, make KSHV a powerful model system for studying the biological roles of putative ORFs. Small is not necessarily simple. In the words of Andy Warhol, “You need to let the little things that would ordinarily bore you suddenly thrill you”.
Researchers

Kevin Brackett
Kevin is investigating the virus-host interactions of Kaposi's Sarcoma-associated Herpesvirus via genetic engineering with CRISPRi technology.

Roy Garcia
Roy is studying the function of KSHV's novel viral small open reading frame PAN 1.1
He is currently a PhD candidate at UT Southwestern Medical Center

Michelle Grunberg
Michelle will it optimizing a Cas9 based system for virus mutagenesis in order to investigate the function of novel KSHV smORFs.
She is a research specialist in the Myhrvold Lab at Princeton University Molecular Biology

Colleen Mispagel
Colleen worked on optimizing a CRISPR-Cas9 based method for the mutagenesis of KSHV's viral genome and is also investigating the effects of Zika virus exposure on the eye during development. She will be applying to medical school.

Ameera Mungale
Ameera is an Arias Lab alumna who previously worked on developing a CRISPRi system for silencing KSHV genes in different host cell lines. She is now a researcher at NIH in the Medical Genetics and Ophthalmic Genomics Unit
She is currently a PhD Student at Georgetown University
