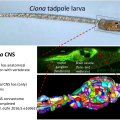
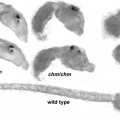
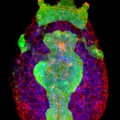
A comprehensive understanding of nervous system function requires integration of multiple types of data, from behavioral to anatomical. We use an organism with unique attributes for multimodal nervous system study: the simple chordate, Ciona robusta. The Ciona larval tadpole CNS shows strong conservation with those of vertebrates, including homologs of the forebrain, midbrain/hindbrain, and spinal cord. Despite these conserved vertebrate-like features the Ciona larval tadpole is barely 1 mm long, and the CNS has only 177 neurons. While the development of the Ciona CNS has been the subject of investigation for many years, new research opportunities have opened with the recent completion of the Ciona tadpole CNS connectome. Our goal is to build from the connectome data to identify correlates of behavior at the level of single, identifiable neurons. The larvae rely on visual, geotactic, tactile and possibly chemosensory inputs to navigate, detect substrates, and perhaps avoid predation. Our studies have focused on the negative phototaxy behavior of Ciona larvae. Larvae orient themselves in a light field using short bursts of spontaneous helical swimming, which then may be followed by sustained swim away from the light source. We have developed models for negative phototaxy taking into account the several photoreceptor groups and their connectivity. In addition, the transparency and small size of Ciona tadpoles make them ideal for live imaging of brain activity using genetically encoded Ca2+ and voltage indicators. We have collected GCaMP recordings from agarose-immobilized tadpoles showing both spontaneous and dimming-induced behaviors using a sheet plane illumination microscope. From the dataset we have been able identified individual cells activated. In summary the connectome makes specific and novel predictions about patterns of neural activity, and the relative simplicity of the Ciona CNS will provide an innovative system for validating such predictions.